I’ve posted up a collection of functions to read from various gridcell and station archives into organized time series objects here. Read functions are included for HadCRUT3, HAdCRUT2, GISS gridded, GHCN v2, GISS station, GSN, meteo.ru, unaami and G02141. I’ll probably add to this from time to time now that I’ve figured out an organization for this. These read functions enable some pretty concise programming as shown below. Thanks to Nicholas for resolving problems with reading zip files and the weird GISS set up, which really unblocked an exercise that had annoyed me for some time.
In general, the read functions will return an R-object (list) with three components $raw; $anom and $normal. For example,
station.ghcn< -read.ghcnv2(id0=29312)
is sufficient to return first – raw – the monthly time series of raw values; normal – the monthly 1961-1990 normals using GHCN as default if insufficient values to calculate within the specified set; anom – the monthly anomaly series.
In some cases, there are more than 1 version for a WMO station number. In such cases, all series are returned together with an average which is named “avg”. I’ve included a function ts.annavg to convert a univariate time series to an annual average with a parameter (default M=6) for a minimum to take an annual average.
station.ghcn.ann< – ts.annavg(station.ghcn$anom[,"avg"])
I’ve posted up a short scripts to show these functions can be used to collate 7 series for studying the Barabinsk gridcell. I’ve shown it here in WordPress code, which is often problematic, so look at the ASCII version if you want to try it. Basically, each line will recover the data from a different archive (provided that some prior unzipping has been done as described below). So it’s a pretty concise way of organizing the data versions (all keyed here merely through the WMO identification).
library(ncdf)
id0< -29612
source("http://data.climateaudit.org/scripts/gridcell/collation.functions.txt")
source("http://data.climateaudit.org/scripts/gridcell/load.stations.txt")
info<-stations[stations$wmo==id0,];info
# id site wmo version long pop lat
#1771 222296120000 BARABINSK 29612 0 78.37 37 55.33
#GRIDDED VERSIONS
station.hadcru3<-read.hadcru3(lat=info$lat,long=info$long) #monthly
station.hadcru2<-read.hadcru2(lat=info$lat,long=info$long) #monthly
station.gissgrid<-read.giss.grid(lat=info$lat,long=info$long)
#STATION VERSIONS
station.ghcn<-read.ghcnv2(id0)
#Error in try(dim(v2d)) : object "v2d" not found
#but proceeds to load
station.normal<-station.ghcn[[3]]
station.giss<-download_giss_data(id0)
station.gsn<-read.gsn(id0)
station.meteo<-read.meteo(id0)
combine<-ts.union(ts.annavg(station.hadcru3),ts.annavg(station.hadcru2),station.gissgrid,ts.annavg(station.ghcn$anom[,"avg"]),
ts.annavg(station.giss$anom[,"avg"]),ts.annavg(station.gsn$anom),ts.annavg(station.meteo$anom) )
dimnames(combine)[[2]]<- c("hadcru3","hadcru2","gissgrid","ghcn2","giss","gsn","meteo")
combine<-ts(combine[(1881:2006)-1849,],start=1881)
par(mar=c(3,3,1,1))
ts.plot(combine[,1:7],col=1:7)#,lwd=c(2,2,1,1,1,1,1),xlab="",ylab="")
legend(1875,2.6,fill=1:7,legend=c("Hadcru3","HadCRU2","GISS Grid","GHCN2","GISS","GSN","Meteo.ru"))
This yields this following spaghetti graph on my machine:
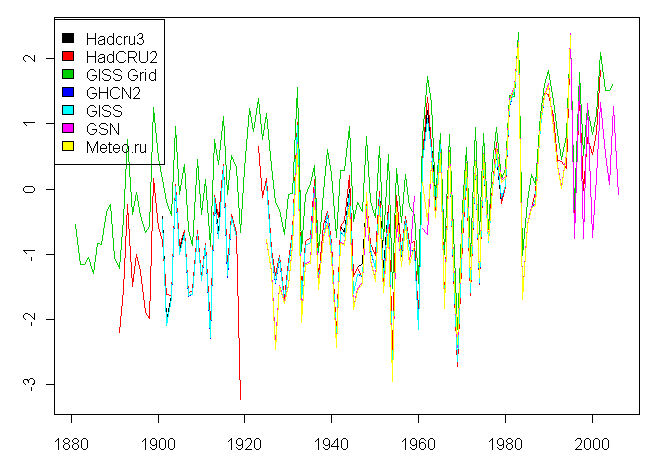
Now you’ll have to have installed the ncdf package and to have downloaded three large zipped files. I’ve not tried to include the unzipping of these large files in the programming since it’s probably a get idea to do this manually to be sure that you get it right. The three large files are:
HadCRUT3 – downloaded and unzipped from http://hadobs.metoffice.com/hadcrut3/data/HadCRUT3.nc into “d:/climate/data/jones/hadcrut3/HadCRUT3.nc” #5×5 monthly gridded from CRU
GISS Gridded – downloaded and unzipped from ftp://data.giss.nasa.gov/pub/gistemp/netcdf/calannual-1880-2005.nc into “d:/climate/data/jones/giss/calannual-1880-2005.nc” #2×2 degree annual gridded from GISS
GHCN Station data – # http://www1.ncdc.noaa.gov/pub/data/ghcn/v2/v2.mean.Z #unzipped, read and re-collated into “d:/climate/data/jones/ghcn/v2d.tab” #see this script for the re-collation to the R-object
HadCRUT2 – downloaded from HAdCRU and re-collated into “d:/climate/data/jones/hadcrut2/hadcruv.tab” – I posted up a script to do this some time ago, but this probably needs to be reviewed.
There may very well be some residual references to things on my computer that will need to be ironed out to make the routines fully objective. Let me know if you run into problems.
In the mean-time, you should be able to get useful components working and I hope that this helps people to wade through the gridcell data.




